This is achieved by using pairwise losses, e.g. Additionally, metric learning approaches were employed to minimize intra-class variations and maximize inter-class variation. In some cases, additional intra-class samples were generated by geometric transformations 12 or from a separate model 16. Since no external class labels are available, the models learn the representation by performing some pretext tasks, such as to maintain spatial coherence 12, 13 or image contour 16. These methods try to find a discriminative pixel embedding that can be linked to the segmentation classes. More recently, methods for unsupervised semantic segmentation started to emerge 12, 13, 14, 15, 16, 17. 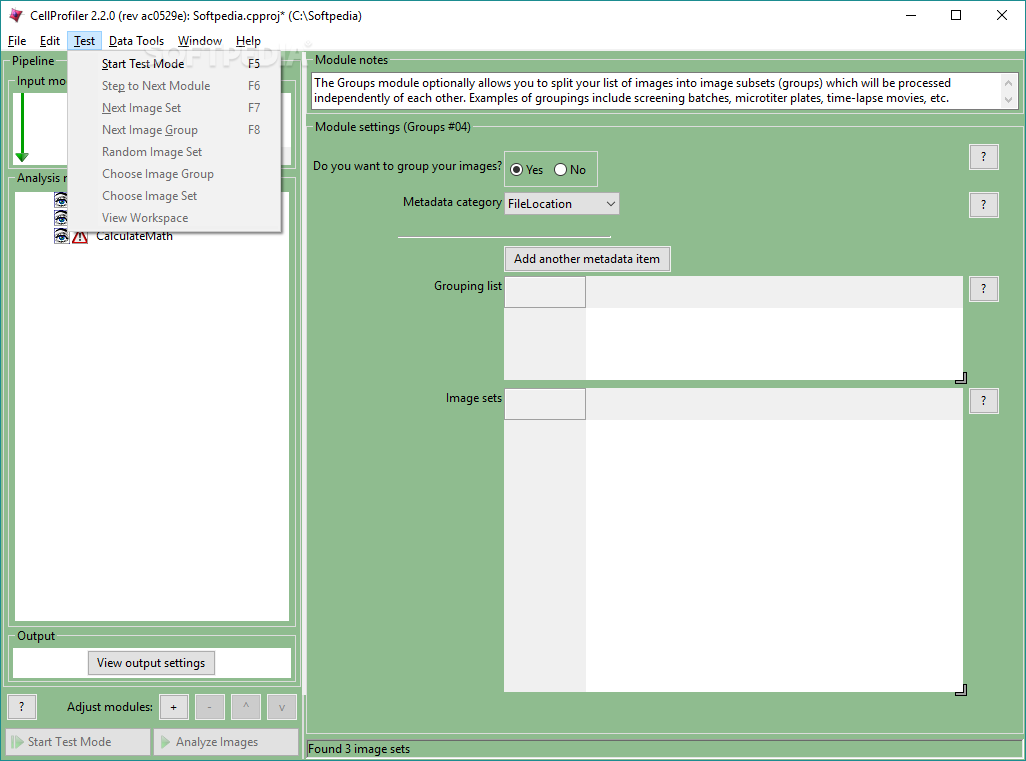
However, high quality annotations of this type are scarce in general. However, to train a CNN for the segmentation task, one typically needs a significant amount of manually labeled training images, in which cell areas and/or cell boundaries are marked by human operators. In particular, various networks with an encoder-decoder architecture (e.g., UNet) have achieved remarkable pixel-level accuracy in the segmentation of objects, including biological cells 7, 8, 9, 10, 11. Therefore, there is a need for more generic, “turn-key” solutions for this task.Ĭonvolutional neural networks (CNN) have been shown to be highly efficient on various kinds of image processing tasks, including semantic and instance segmentations 3, 4, 5, 6. changing the objective) often requires a redesign and/or reoptimization of the segmentation algorithm. Switching to a different cell type (with different morphological features), changing imaging modality (e.g., from epi-fluorescence to confocal), or even imaging settings (e.g. For example, an algorithm that is well-optimized for a specific membrane fluorescent marker will not work on images of a different fluorescent marker, nor on bright field images. Unfortunately manual feature definitions are usually highly context-specific and require task-dependent tuning to work well. Traditional approaches to cell segmentation rely on manually-crafted feature definitions that allow the algorithmic recognition of cellular area and cell border 1, 2. We also post guided exercises as part of our educational outreach effort.Automated cellular segmentation from optical microscopy images is a critical task in many biological researches that rely on single-cell analysis. Simple nuclei identification tutorial ( sample data) (courtesy of the German BioImaging network) Performing a colocalization assay ( relevant example pipeline) Using the Worm Toolbox for image analysis of C. Identifying and measuring cells: Cytoplasm-nucleus translocation assay ( relevant example pipeline)Ĭalculating and applying illumination correction for images ( relevant example pipeline) Identifying, measuring, and classifying yeast colonies ( relevant example pipeline) Using the Input modules in CellProfiler 2.1: Using CellProfiler for Quantitative Image Analysis The NIH has published a introductory chapter of “best practices” for image-based high-content screening (in which CellProfiler is mentioned) as part of the Assay Guidance Manual, and our group has published a more advanced follow-up chapter on image analysis methods. Our introduction to automated image analysis principles and practicalities is published as an educational article at PLoS. Technical descriptions of CellProfiler and CellProfiler Analyst software can be found in our papers while more written tutorials can be found on the CellProfiler GitHub page. Visit our YouTube playlist for video tutorials on CellProfiler, CellProfiler Analyst, segmentation strategies, how to construct pipelines, and much more. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
